A team of scientists isolated 13 immune system compounds, called nanobodies, which show promise for treating COVID-19. The nanobodies, isolated from a llama’s immune cells, blocked the SARS-CoV-2 virus from entering human cells. Negin Martin, Ph.D., director of the NIEHS Viral Vector Core, contributed to the research. The study was posted Aug. 23 on Cold Spring Harbor Laboratory’s bioRxiv preprint server.
Scientists previously learned that the novel coronavirus enters human cells by latching on to a protein on a cell’s surface known as the ACE-2 receptor. That receptor is so named because it binds to an enzyme called ACE-2, the way a lock and key fit together. ACE-2 helps regulate blood pressure, inflammation, and other processes. Interestingly, the coronavirus that caused the 2002 SARS outbreak uses the same protein.
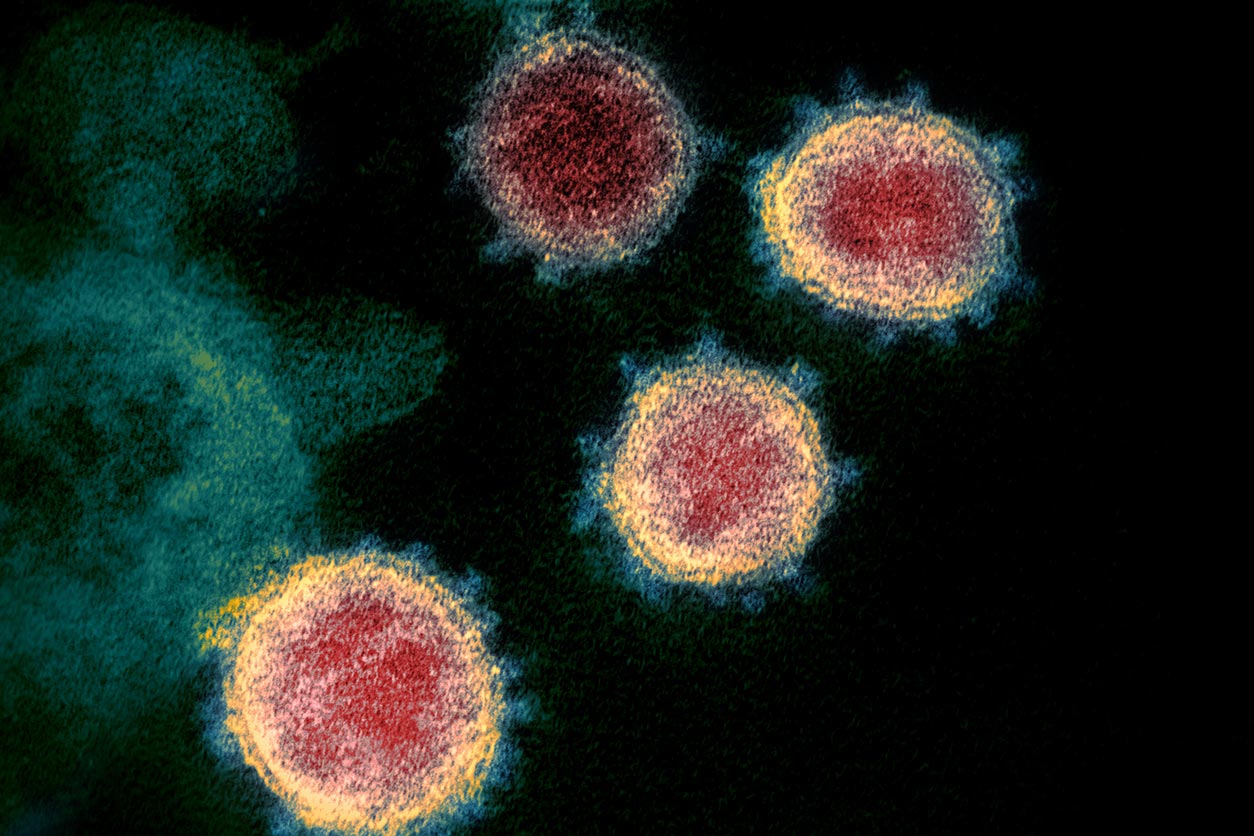 SARS-CoV-2 virus particles emerging from cells. NIH-CoVnb-112 binds with the spike protein, blocking the virus from gaining entry to a new cell. (Photo courtesy of National Institute of Allergy and Infectious Disease)
SARS-CoV-2 virus particles emerging from cells. NIH-CoVnb-112 binds with the spike protein, blocking the virus from gaining entry to a new cell. (Photo courtesy of National Institute of Allergy and Infectious Disease)Preprint caveat
In the context of a global pandemic, many researchers are publishing their results on preprint servers to speed development of treatments and vaccines. Such findings have not been rigorously vetted by experts through the usual peer review process, although most authors also submit their papers to peer-reviewed journals at the same time.
The new study's senior author, David Brody, M.D., Ph.D., and his team made the same choice. Brody is with the National Institute of Neurological Disease and Stroke (NINDS) and directs the Center for Neuroscience and Regenerative Medicine (CNRM) at the Uniformed Services University of the Health Sciences. CNRM is a joint venture with NINDS focused on traumatic brain injury. First author Thomas “TJ” Esparza is with the Henry Jackson Foundation for the Advancement of Military Medicine as well as NINDS.
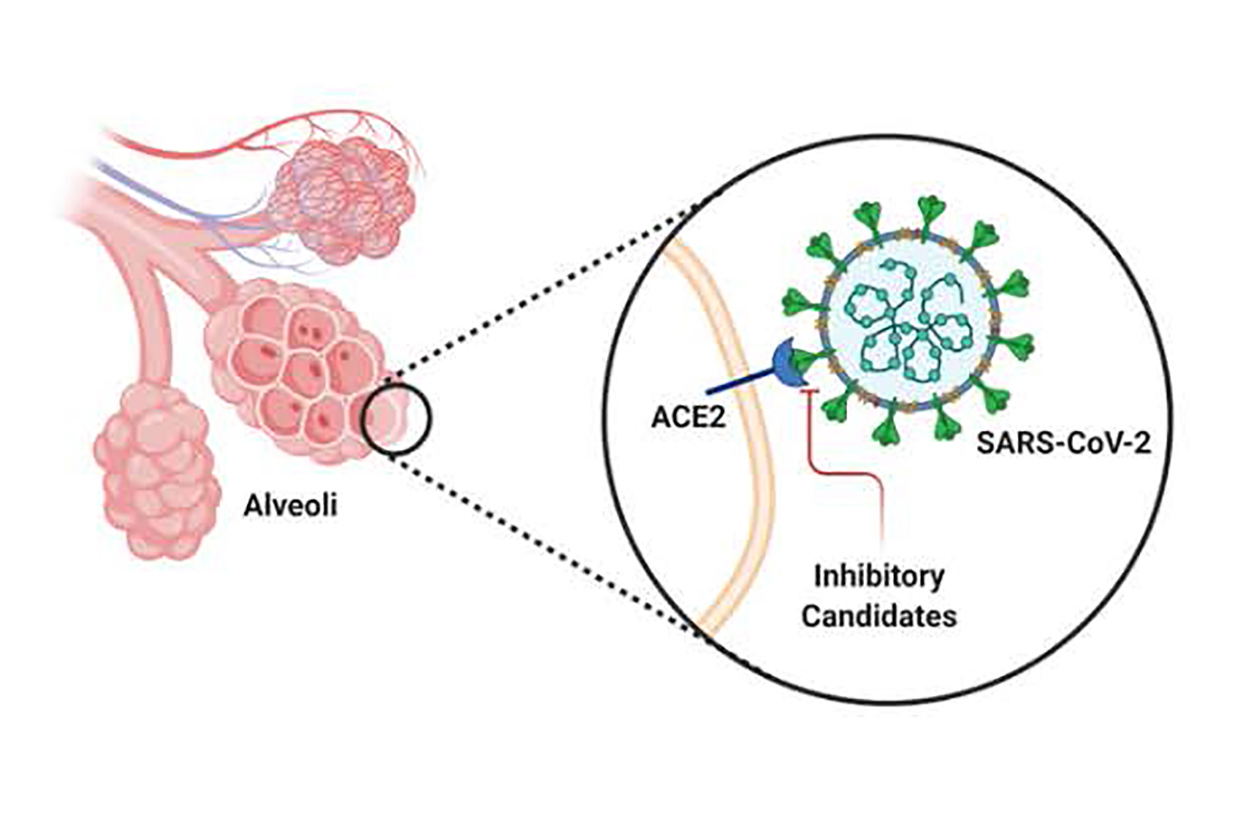 Illustration of the SARS-CoV-2 spike protein, with receptor binding domain in contact with the human ACE-2 receptor on the surface of a lung epithelial cell. (Photo courtesy of TJ Esparza)
Illustration of the SARS-CoV-2 spike protein, with receptor binding domain in contact with the human ACE-2 receptor on the surface of a lung epithelial cell. (Photo courtesy of TJ Esparza)Nano tool
Nanobodies are antibody fragments that can be produced on a large scale at low cost, the study authors wrote. Advanced screening strategies helped the team identify nanobody DNA sequences from a llama inoculated with the novel coronavirus (see sidebar).
 “The lead candidate was a very hardy protein — still very effective after being nebulized,” said Martin. “It is wonderful to be able to support such a strong project through NIH collaborative efforts.” (Photo courtesy of Steve McCaw)
“The lead candidate was a very hardy protein — still very effective after being nebulized,” said Martin. “It is wonderful to be able to support such a strong project through NIH collaborative efforts.” (Photo courtesy of Steve McCaw)Esparza tested the sequences to see which ones bound most strongly to the spike protein. From these, the team isolated 13 that both bound to the SARS-CoV-2 spike protein and effectively blocked its interaction with the ACE-2 receptor. “These sequences were distinct from the previously published sequences that also bind SARS-CoV-2 spike protein,” the authors wrote.
The strongest of the baker’s dozen, which they dubbed NIH-CoVnb-112, proved effective in the human embryonic kidney cell line used in the lab. Martin analyzed the effects of that nanobody on viral entry and infection. Brody and Esparza confirmed efficacy in three genetic variants of the spike protein currently circulating around the world.
Importantly, this lead candidate retained structural integrity and potency after delivery through a nebulizer. The authors suggested that NIH-CoVnb-112 shows promise for treatment, as well as preventative and diagnostic uses. However, further structural and chemical characterization is needed, as well as extensive testing to ensure safety and efficacy.
Pandemic pivot
Like other scientists, Brody and Esparza saw the spike protein–ACE-2 receptor connection as a vulnerable target to stop the spread of SARS-CoV-2 and the pandemic it unleashed. But how do scientists working on traumatic brain injury shift to infectious disease?
 Cormac lives on a farm in Washington, where he contributes to scientific discovery. (Photo courtesy of Triple J Farms)
Cormac lives on a farm in Washington, where he contributes to scientific discovery. (Photo courtesy of Triple J Farms)Enter the llama and its nanobodies. As members of the camelid family — along with alpacas and dromedaries — llamas produce a class of immunoglobulins with an unusual characteristic. The portion of the protein that recognizes foreign substances, called the antigen recognition domain, can be expressed as a fragment called a nanobody.
Esparza and Brody were using llama nanobodies to improve human brain imaging for studies of disease progression and treatment response. “Like many researchers, we were exploring how our methods might contribute to COVID-19 research,” Esparza said. “We felt compelled to serve; it’s part of our ethos.”
Among the challenges was designing a study that needed only one person at a time in the laboratory. As to how they came to collaborate with Martin, “We take our hat off to NIH leadership, who set up a dashboard of resources for COVID-19 projects,” Esparza explained.
“Negin listed herself as able to run the kind of assays we needed,” Brody said. “Her assay would have taken us months to set up. She has been a terrific collaborator.”
Citation: Esparza TJ, Martin NP, Anderson GP, Goldman ER, Brody DL. 2020. High affinity nanobodies block SARS-CoV-2 spike receptor binding domain interaction with human angiotensin converting enzyme. bioRxiv; doi:10.1101/2020.07.24.219857 [Online 23 August 2020].