The 2021 Dr. Martin Rodbell Lecture Series Seminar was a virtual event and featured Aviv Regev, Ph.D., Genentech executive vice president of research and early development (see first sidebar). She delivered her talk, 'Cell Atlases as Roadmaps to Understand and Treat Disease,' on April 13. Benedict Anchang, Ph.D., a Stadtman Investigator in the NIEHS Biostatistics and Computational Biology Branch, hosted the seminar.
'Dr. Aviv’s research revolutionizes our understanding of biology and disease at a systems level, which is key to advancing the field of precision medicine,' Anchang said.
NIEHS Director Rick Woychik, Ph.D., began the session by honoring the memory and scientific contributions of former NIEHS Scientific Director Martin Rodbell, Ph.D. He shared the 1994 Nobel Prize in Physiology or Medicine with Alfred Gilman, M.D., Ph.D.
 Regev presented work she conducted in her lab when she was a Howard Hughes Medical Institute Investigator at MIT and the Broad Institute. She is transitioning her lab to Genentech. (Photo courtesy of Casey Atkins / Broad Institute)
Regev presented work she conducted in her lab when she was a Howard Hughes Medical Institute Investigator at MIT and the Broad Institute. She is transitioning her lab to Genentech. (Photo courtesy of Casey Atkins / Broad Institute)Correlation between computer science and biology
Regev said that in the 1960s, Rodbell illuminated the similarities between the information processing systems of computers and biological organisms. For instance, Rodbell used to say cell receptors were discriminators that receive input and act like classifiers. Transducers process this input or information across the cell membrane, whereas amplifiers intensify these signals to initiate cellular reactions or transmit information to other cells.
'He had a cell-centric view, but also a computational view,' Regev said.
Other seminar attendees also saw the interconnection between Regev’s and Rodbell’s work. 'During her outstanding lecture I was struck by how excited Dr. Rodbell would likely have been by the power of single cell technologies,' said Hans Luecke, Ph.D., NIEHS associate scientific director. 'Dr. Regev clearly shares his sense of curiosity.'
Dynamics of differentiation
Regev’s research focus is single cell genomics, the study of individual cells using genetic tools. Over the past several years, the methods she and others have developed have shed light on some of the most fundamental aspects of cells and tissues. One of those areas is cell fate.
She explained that the developmental trajectory from a stem cell to its differentiated form —whether a neuron, muscle, or skin cell — can be monitored experimentally along a time course. Several years ago, in collaboration with members of the laboratory led by Alexander Schier, Ph.D., she and her group profiled zebrafish cells at one-hour intervals for 12 time points during early development.
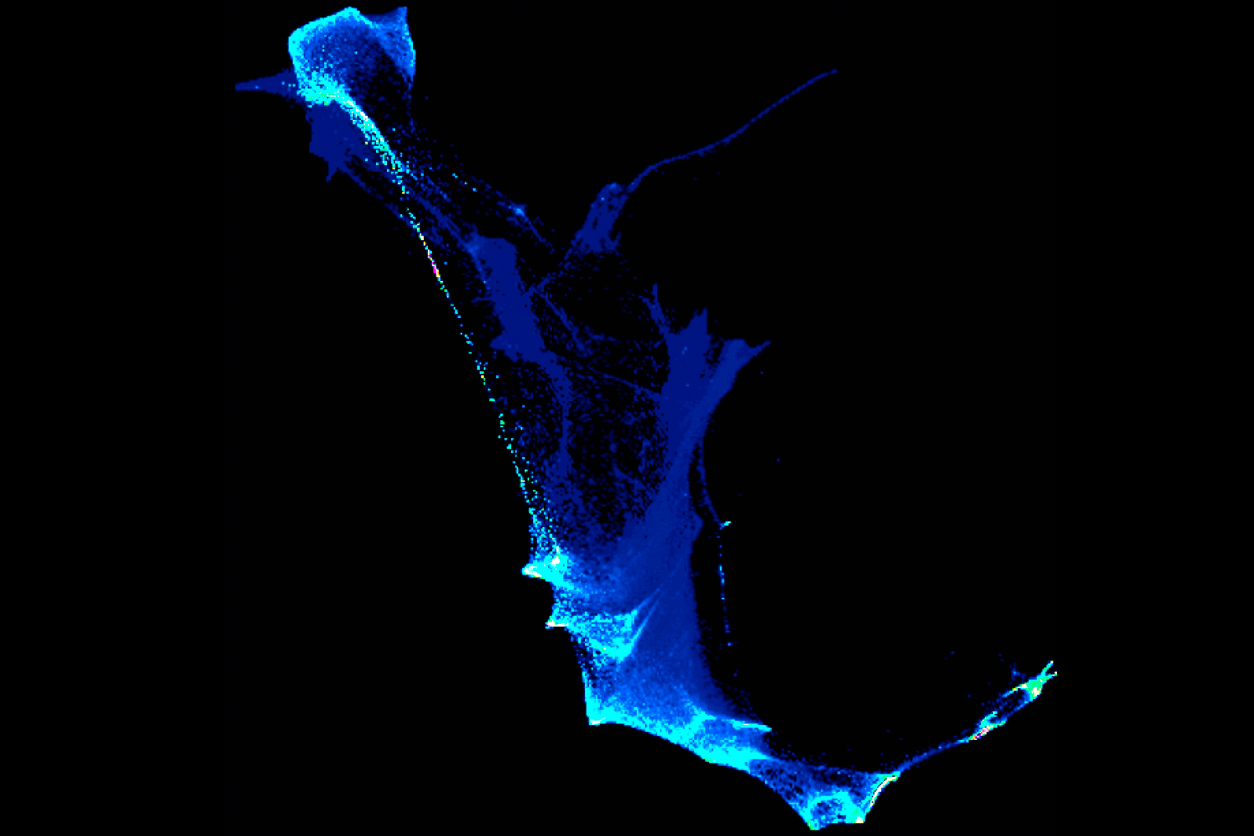 This cell lineage map is compiled from a video depicting zebrafish cell differentiation during early development. The bright sections represent cell ancestors. (Image courtesy of Aviv Regev)
This cell lineage map is compiled from a video depicting zebrafish cell differentiation during early development. The bright sections represent cell ancestors. (Image courtesy of Aviv Regev)With computational analysis, the cell profiles could be organized in two-dimensional space. Dozens of cell types were produced after 12 hours. Cells were organized both by the progression of time and how they diversified. When the cells were color coded by time point, the colors blended due to some asynchrony.
 Anchang is an Earl Stadtman Investigator in the NIEHS Biostatistics and Computational Biology Branch where he leads the Computational and Systems Biology Group. (Photo courtesy of Steve McCaw / NIEHS)
Anchang is an Earl Stadtman Investigator in the NIEHS Biostatistics and Computational Biology Branch where he leads the Computational and Systems Biology Group. (Photo courtesy of Steve McCaw / NIEHS)To understand how the cells differentiated into various types, Regev and her colleagues — especially Jeff Farrell, Ph.D., then a postdoc — developed an algorithm called URD to reconstruct the process (see second sidebar). The tree that URD produced allowed them to see gene progression across every cellular lineage. Within each path, they could see active genes.
'This shows how a cell’s profile carried information about its developmental potential,' Regev said. 'Using URD, we can play the tape backward to trace ancestors or forward to trace descendants.'
Regev also discussed other aspects of her research, including combining spatial and differentiation maps, cell lineages, and causal mechanisms in disease. She saved the last part of her talk for her favorite initiative.
The Human Cell Atlas
According to Regev, cell maps are useful, so why not have an atlas of human cells that combines those maps? The Human Cell Atlas is an international community of scientists working to create comprehensive reference maps of all human cells.
Because cells are the fundamental units of life, the atlas will be the basis for both understanding human health and diagnosing, monitoring, and treating disease. In all her seminars, Regev encourages scientists to join the endeavor.
'You don’t need a grant,' Regev said, 'just go to Human Cell Atlas and contribute to the effort.'
Citation: Farrell JA, Wang Y, Riesenfeld SJ, Shekhar K, Regev A, Schier AF. 2018. Single-cell reconstruction of developmental trajectories during zebrafish embryogenesis. Science 360(6392):eaar3131.









