Our environment can affect our health at the most fundamental level, influencing our genome by changing how our genes are read or expressed. This concept — known as environmental epigenetics — has transformed the way that scientists look for links between environmental exposures and disease. For example, Harvard researcher Bernardo Lemos, Ph.D., has focused his research on a single organelle that sits within the cell’s nucleus called the nucleolus.
 Lemos is an associate professor of environmental epigenetics at the Harvard T.H. Chan School of Public Health. (Photo courtesy of Bernardo Lemos)
Lemos is an associate professor of environmental epigenetics at the Harvard T.H. Chan School of Public Health. (Photo courtesy of Bernardo Lemos)During his Mar. 30 NIEHS Keystone Science Lecture, Lemos explained how the study of this “tiny nucleus” is yielding insights into aging, carcinogenesis, and chronic diseases.
“It’s a really rich organelle,” he said. “And from a basic biology perspective, it’s where the inspiration for my lab resides.”
Tiny nucleus
The nucleolus is made up of an array of genomic segments known as ribosomal DNA (rDNA), which encode the machinery needed to synthesize all the body’s proteins. Lemos believes that rDNA is particularly sensitive to the environment and thus could provide informative biomarkers of aging and environmental exposures.
As Lemos explained, aging is influenced by the environment and is among the most important known risk factors for predicting disease and mortality. Multiple studies have shown that aging — and the chronic diseases that come with it — can be accelerated by exposure to environmental toxicants and slowed by a relatively simple intervention: cutting calories by 20%.
“As we learn more about environmental exposures, we will likely find how exposures drive aging in the opposite direction, that is, accelerate aging,” Lemos said.
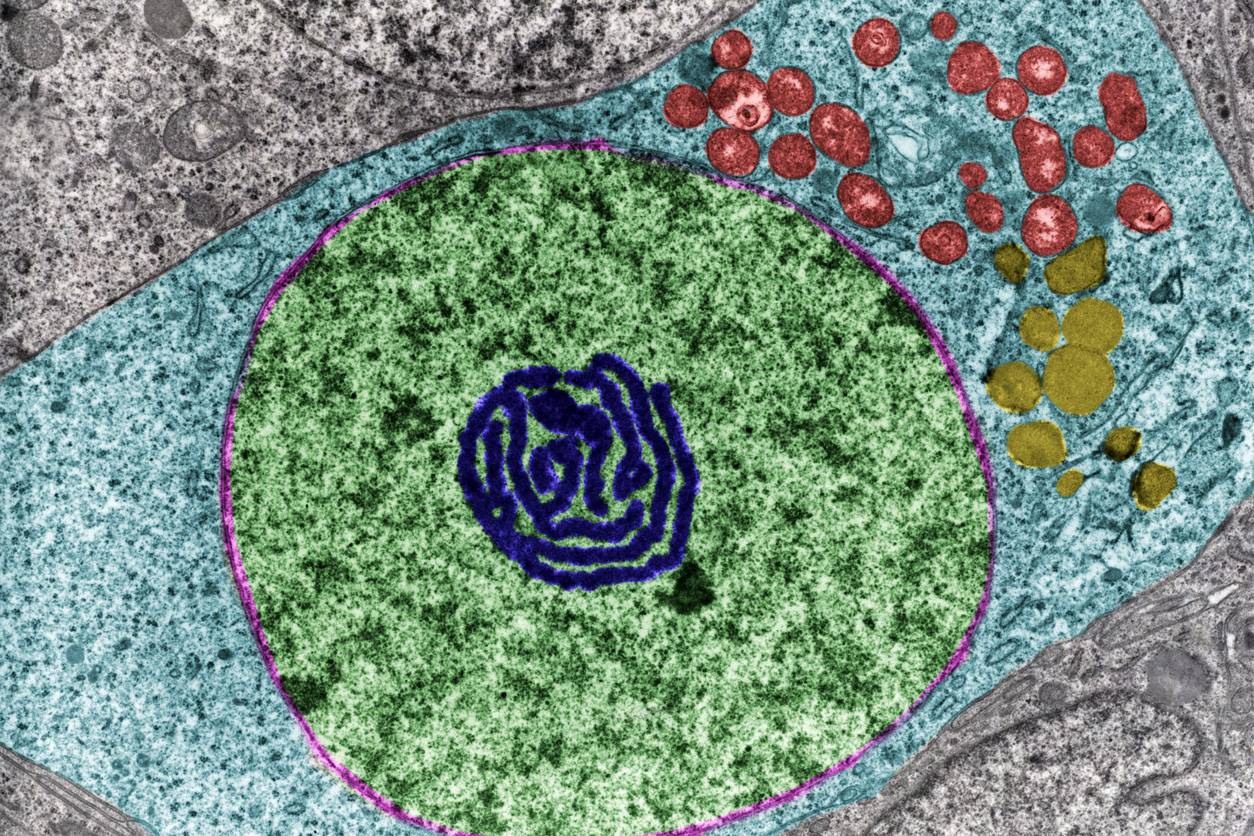 A false color transmission electron microscope image showing the cell’s nucleus (green) and nucleolus (dark blue), a structure made up of rDNA. (Photo courtesy of Jose Luis Calvo / Shutterstock)
A false color transmission electron microscope image showing the cell’s nucleus (green) and nucleolus (dark blue), a structure made up of rDNA. (Photo courtesy of Jose Luis Calvo / Shutterstock)His lab discovered that specific methylation patterns in rDNA can serve as an evolutionarily conserved “clock” of biological aging. Lemos showed that this clock was conserved between mice, other mammals, and humans. Interestingly, he found that this epigenetic clock was responsive to changes in the environment. When his team assessed mice and rats on a calorie restricted diet, the rodents’ clocks slowed down.
Heavy metals
Lemos is currently funded by NIEHS to study how exposure to a specific environmental chemical called hexavalent chromium affects rDNA. Hexavalent chromium is a form of the metallic element chromium that has been shown to cause lung cancer. As part of his study, Lemos exposed bronchial epithelial cells — which act as the interface between the external environment and the lungs — to various doses of hexavalent chromium for 24 hours. Then he tracked changes in genome-wide gene expression for seven days.
 Shaughnessy said Lemos’s research translates basic scientific understanding into public health with the goal of characterizing the susceptibility of individuals to the effects of environmental exposures. (Photo courtesy of Steve McCaw / NIEHS)
Shaughnessy said Lemos’s research translates basic scientific understanding into public health with the goal of characterizing the susceptibility of individuals to the effects of environmental exposures. (Photo courtesy of Steve McCaw / NIEHS)He found that the environmental exposure affected the nucleolus and triggered widespread and persistent expression and DNA methylation changes in thousands of genes.
“Where do we think that responsiveness of gene expression is coming from?” Lemos said. “We think, fundamentally, it is coming from the organization of the epigenome.”
Lemos was hosted by Daniel Shaughnessy, Ph.D., a health science administrator in the NIEHS Exposure, Response, and Technology Branch.
“Bernardo’s research on the effects of environmental stress on ribosomal DNA, together with his findings on methylation patterns of rDNA genes as a biological clock, is truly innovative,” said Shaughnessy. “[The methylation patterns] have the potential for serving as informative biomarkers of the effects of the environment on aging and disease risk.”
Microplastics
Another exposure that Lemos has studied at the epigenetic level is microplastics, the tiny pieces of plastic now being detected throughout the environment. Lemos explained how his lab has surveyed gene expression in the guts of mice exposed to such material. They discovered changes in the expression of genes involved in a variety of biological processes, including oxidative response, lipid metabolism, immune response, and hormone metabolism.
Lemos said that in the future, he would be interested in investigating the effects of microplastic pollution in specific disadvantaged communities and is open to collaborating with anyone with expertise in that area.
“All of the work that I have presented — even the work that was not directly funded by NIEHS — would not have been possible without the support of the institute,” said Lemos. “For that, I am truly grateful.”
Citations:
Lou J, Yu S, Feng L, Guo X, Wang M, Branco AT, Li T, Lemos B. 2021. Environmentally induced ribosomal DNA (rDNA) instability in human cells and populations exposed to hexavalent chromium [Cr (VI)]. Environ Int 153:106525.
Wang M, Lemos B. 2019. Ribosomal DNA harbors an evolutionarily conserved clock of biological aging. Genome Res 29(3):325−333.
Zhang Y, Wolosker MB, Zhao Y, Ren H, Lemos B. 2020. Exposure to microplastics cause gut damage, locomotor dysfunction, epigenetic silencing, and aggravate cadmium (Cd) toxicity in Drosophila. Sci Total Environ 744:140979.
(Marla Broadfoot, Ph.D., is a contract writer for the NIEHS Office of Communications and Public Liaison.)









