Gary Siuzdak, Ph.D., from the Scripps Research Institute, highlighted exciting technologies that he said will advance the field of metabolomics and a wide range of scientific discovery, during a Dec. 7 NIEHS lecture. Metabolomics is the large-scale study of chemical reactions involving metabolites, which are small molecules that play important roles in cells, tissues, and organisms.
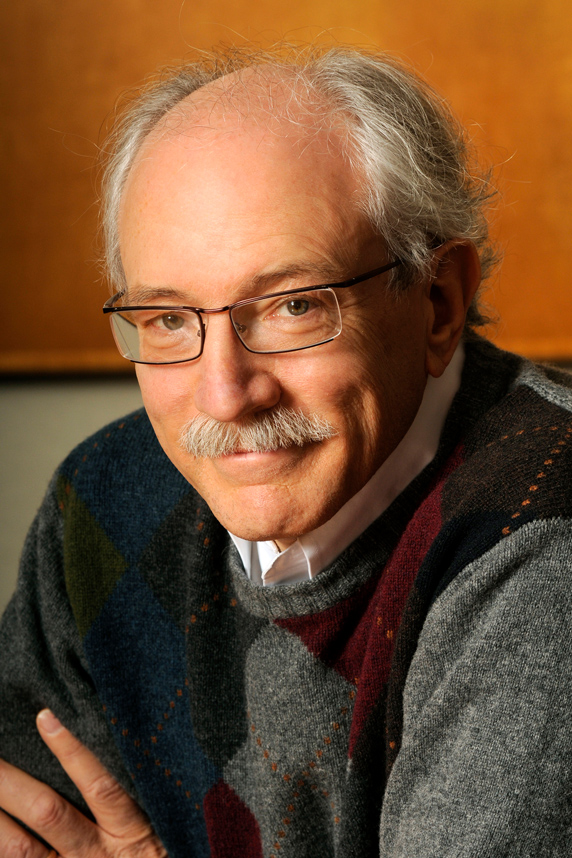
 “We have created unique metabolomics research tools,” said Siuzdak. “There is a lot of interest in making them even more comprehensive.” (Photo courtesy of Gary Siuzdak)
“We have created unique metabolomics research tools,” said Siuzdak. “There is a lot of interest in making them even more comprehensive.” (Photo courtesy of Gary Siuzdak)Metabolites’ role in health and disease
According to Siuzdak, research in this field originally focused on identifying metabolites that serve as biological signs of disease, which scientists call biomarkers. However, metabolomics has evolved into a more comprehensive tool for understanding how metabolites themselves can influence health and illness.
“The most important area where metabolomics can be applied is in looking for active metabolites that affect physiology,” Siuzdak said. “For example, metabolites can impact and even improve the way we respond to medicine or exposure to toxic agents.”
Enabling scientific discovery
Siuzdak developed data analysis platforms — called XCMS and METLIN — that enable scientists to discover how metabolites can alter critical biological processes, and the tools have been cited in more than 10,000 scientific projects, he noted.
“Through XCMS and METLIN, which now contains detailed data on 860,000 molecular standards, the Scripps Center for Metabolomics has strengthened research worldwide, across a variety of disciplines,” said Siuzdak, the center’s director.
“Continued development of databases like METLIN is vital to success of the metabolomics field,” noted David Crizer, Ph.D., a chemist in the NIEHS Division of the National Toxicology Program. He is a member of the institute’s Metabolomics Cross-Divisional Group, which hosted Siuzdak’s talk (see sidebar).
 “This seminar was a great opportunity to learn about things to consider when designing metabolomics studies,” Crizer said. (Photo courtesy of Steve McCaw / NIEHS)
“This seminar was a great opportunity to learn about things to consider when designing metabolomics studies,” Crizer said. (Photo courtesy of Steve McCaw / NIEHS)Identifying key molecules
METLIN is designed to help scientists identify molecules in organisms, whether metabolites, toxicological agents, or other chemical entities, according to Siuzdak. He noted that the database encompasses more than 350 chemical classes, and there now are more than 50,000 registered users in 132 countries.
“Our goal is to identify as many metabolites and other chemical entities as possible, and given the advances in other fields of biology, this data is long overdue,” Siuzdak said.
“We are finding metabolites that were previously unknown, quite regularly,” he added. “The more comprehensive METLIN is, the better chance we have of eventually identifying all molecules. To this end, I am constantly looking for ways to facilitate growth of the platform.”
Gut check
A metabolite called indole-3-propionic acid (IPA) is of particular interest to Siuzdak. IPA is a human gut–derived metabolite originally identified by his lab in a 2009 paper in the Proceedings of the National Academy of Sciences, and it has since been examined in thousands of studies. Researchers have discovered that it is a multifunctional molecule that can aid immune function, among other roles.
“In retrospect, it makes sense that a metabolite derived from a gut microbe could modulate the immune system, which is probably why it still generates so much excitement,” he said.
IPA could be especially relevant with respect to autoimmune diseases, Siuzdak added.
“For example, most people who die from COVID-19 don’t succumb to the virus but from an overactive immune response that causes them to develop respiratory ailments,” he said. “A metabolite that modulates this effect could be very beneficial,” noted Siuzdak.
“Overall, we are pursuing one primary goal in the development of METLIN, which is to use experimental data generated from molecular standards to help identify these key, physiologically relevant molecules,” he said.
Citation: Wikoff WR, Anfora AT, Liu J, Schultz PG, Lesley SA, Peters EC, Siuzdak G. 2009. Metabolomics analysis reveals large effects of gut microflora on mammalian blood metabolites. Proc Natl Acad Sci U S A 106(10):3698–3703.
(John Yewell is a contract writer for the NIEHS Office of Communications and Public Liaison.)