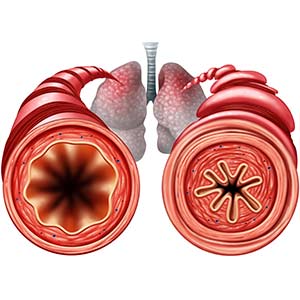
An international team of scientists led by NIEHS identified epigenetic markers — chemical tags attached to DNA — that may indicate a newborn baby’s risk of developing asthma.
The results, published Dec. 21 in the Journal of Allergy and Clinical Immunology, were drawn from data collected by the Pregnancy And Childhood Epigenetics (PACE) consortium.
“This study shows that we can find some potential epigenetic biomarkers of later asthma risk, and that larger studies will likely generate many more hits,” said senior study author Stephanie London, M.D, Dr.P.H., head of the NIEHS Genetics, Environment, and Respiratory Disease Group.
“Epigenetic variation, combined with genetic sequence variation, might more reliably identify at birth the children who are going to develop asthma. This could inform interventions to prevent asthma,” she explained.
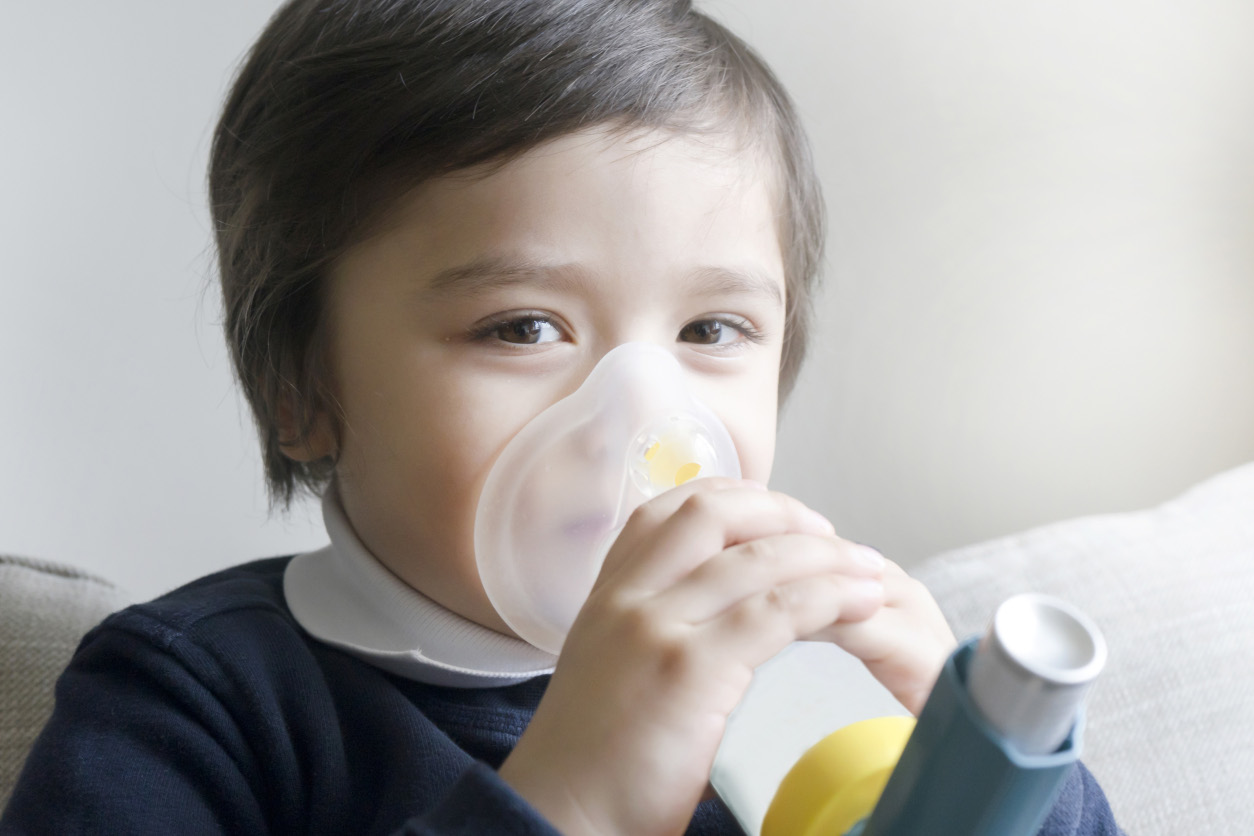 Children who are at risk of developing asthma may be identified by the new approach reported by London and colleagues.
Children who are at risk of developing asthma may be identified by the new approach reported by London and colleagues.Breaking the code
Asthma is the most common chronic disease of childhood, affecting one in 12 children in the United States. Large-scale scans of variations in the DNA sequence — known as genome-wide association studies — have found many genetic variants related to asthma. However, these variants only explain a portion of disease risk.
Scientists began looking beyond the DNA sequence for a better understanding of asthma. Recently, they turned to epigenetics, which is the study of the chemicals and proteins that bind to DNA to activate or silence genes without changing the underlying sequence.
Changes linked to asthma development
 London is a leader in the PACE consortium, an international group of researchers using epigenetics to study how environmental exposures in early life affect human disease. (Photo courtesy of Steve McCaw)
London is a leader in the PACE consortium, an international group of researchers using epigenetics to study how environmental exposures in early life affect human disease. (Photo courtesy of Steve McCaw)In this study, London and her colleagues in the PACE consortium focused on the best studied epigenetic modification — methylation, or the addition of methyl tags to DNA. The researchers examined methylation patterns at birth and followed the children until they reached school age to see who developed asthma.
They discovered seven different spots in the genome where methylation rates were higher in children who became asthmatic compared with those who did not.
In a separate meta-analysis, the researchers examined methylation in a different group of children when they were diagnosed. The researchers identified far more spots — 173 in the genome — that exhibited different methylation patterns in relation to asthma. They believe these associations could reflect both the processes that drive the development of asthma and the effects of having the disease.
Druggable targets
In addition to identifying potential biomarkers of asthma, the new findings may shed light on the origins and development of the disease. Many of the methylated regions uncovered in this study include genes that manage immune responses that could cause or worsen asthma. In addition, several of these genes are targets for either approved or investigational drugs.
The researchers plan to continue the study, with the hope that their findings could one day lead to new ways to identify children who are most at risk of developing asthma and to inform new therapies to reduce that risk.
“We expect to revisit this analysis in a few years with an even larger sample size, which may enable us to see differential methylation patterns that are more predictive of the risk of developing asthma,” London said. “We have three times the studies now in the PACE consortium compared with when we started this project.”
Citation: Reese SE, Xu C, den Dekker HT, Lee MK, Sikdar S, Ruiz-Arenas C, Merid SK, Rezwan FI, Page CM, Ulemar V, Melton PE, Oh SS, Yang IV, Burrows K, Soderhall C, Jima DD, Gao L, Arathimos R, Kupers LK, Wielscher M, Rzehak P, Lahti J, Laprise C, Madore AM, Ward J, Bennett BD, Wang T, Bell DA; BIOS consortium, Vonk JM, Haberg SE, Zhao S, Karlsson R, Hollams E, Hu D, Richards AJ, Bergstrom A, Sharp GC, Felix JF, Bustamante M, Gruzieva O, Maguire RL, Gilliland F, Baiz N, Nohr EA, Corpeleijn E, Sebert S, Karmaus W, Grote V, Kajantie E, Magnus MC, Ortqvist AK, Eng C, Liu AH, Kull I, Jaddoe VWV, Sunyer J, Kere J, Hoyo C, Annesi-Maesano I, Arshad SH, Koletzko B, Brunekreef B, Binder EB, Raikkonen K, Reischl E, Holloway JW, Jarvelin MR, Snieder H, Kazmi N, Breton CV, Murphy SK, Pershagen G, Anto JM, Relton CL, Schwartz DA, Burchard EG, Huang RC, Nystad W, Almqvist C, Henderson AJ, Melen E, Duijts L, Koppelman GH, London SJ. 2018. Epigenome-wide meta-analysis of DNA methylation and childhood asthma. J Allergy Clin Immunol; doi: 10.1016/j.jaci.2018.11.043 [Online 21 December 2019].
(Marla Broadfoot, Ph.D., is a contract writer for the NIEHS Office of Communications and Public Liaison.)